Home » Posts tagged '#ACSSanFran'
Tag Archives: #ACSSanFran
PF 06821497
PF 06821497
Cas 1844849-11-1
Designed to treat lymphoma
1(2H)-Isoquinolinone, 5,8-dichloro-2-[(1,2-dihydro-4-methoxy-6-methyl-2-oxo-3-pyridinyl)methyl]-3,4-dihydro-7-[(S)-methoxy-3-oxetanylmethyl]-
MF C22 H24 Cl2 N2 O5,
MW 467.34
PF 06821497
5,8-Dichloro-2-[(4-methoxy-6-methyl-2-oxo-1,2-dihydro-3-pyridinyl)methyl]-7-[methoxy(3-oxetanyl)methyl]-3,4-dihydro-1(2H)-isoquinolinone
1(2H)-Isoquinolinone, 5,8-dichloro-2-[(1,2-dihydro-4-methoxy-6-methyl-2-oxo-3-pyridinyl)methyl]-3,4-dihydro-7-(methoxy-3-oxetanylmethyl)-
- Molecular Formula C22H24Cl2N2O5
- Average mass 467.342 Da
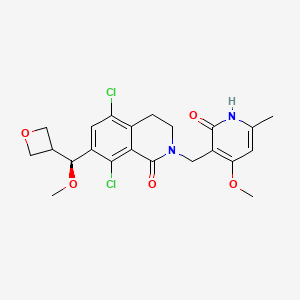
5,8-dichloro-2-[(4-methoxy-6-methyl-2-oxo-1H-pyridin-3-yl)methyl]-7-[(S)-methoxy(oxetan-3-yl)methyl]-3,4-dihydroisoquinolin-1-one
| Inventors | Michael Raymond Collins, Robert Steven Kania, Robert Arnold Kumpf, Pei-Pei Kung, Daniel Tyler Richter, Scott Channing Sutton, Martin James Wythes |
| Original Assignee | Pfizer Inc. |
-
Epigenetic alterations play an important role in the regulation of cellular processes, including cell proliferation, cell differentiation and cell survival. The epigenetic silencing of tumor suppressor genes and activation of oncogenes may occur through alteration of CpG island methylation patterns, histone modification, and dysregulation of DNA binding protein. Polycomb genes are a set of epigenetic effectors. EZH2 (enhancer of zeste homolog 2) is the catalytic component of the Polycomb Repressor Complex 2 (PRC2), a conserved multi-subunit complex that represses gene transcription by methylating lysine 27 on Histone H3 (H3K27). EZH2 plans a key role in regulating gene expression patterns that regulate cell fate decisions, such as differentiation and self-renewal. EZH2 is overexpressed in certain cancer cells, where it has been linked to cell proliferation, cell invasion, chemoresistance and metastasis.
-
High EZH2 expression has been correlated with poor prognosis, high grade, and high stage in several cancer types, including breast, colorectal, endometrial, gastric, liver, kidney, lung, melanoma, ovarian, pancreatic, prostate, and bladder cancers. See Crea et al., Crit. Rev. Oncol. Hematol. 2012, 83:184-193, and references cited therein; see also Kleer et al., Proc. Natl. Acad. Sci. USA 2003, 100:11606-11; Mimori et al., Eur. J. Surg. Oncol. 2005, 31:376-80; Bachmann et al., J. Clin. Oncol. 2006, 24:268-273; Matsukawa et al., Cancer Sci. 2006, 97:484-491; Sasaki et al. Lab. Invest. 2008, 88:873-882; Sudo et al., Br. J. Cancer 2005, 92(9):1754-1758; Breuer et al., Neoplasia 2004, 6:736-43; Lu et al., Cancer Res. 2007, 67:1757-1768; Ougolkov et al., Clin. Cancer Res. 2008, 14:6790-6796; Varambally et al., Nature 2002, 419:624-629; Wagener et al., Int. J. Cancer 2008, 123:1545-1550; and Weikert et al., Int. J. Mol. Med. 2005, 16:349-353.Recurring somatic mutations in EZH2 have been identified in diffuse large B-cell lymphoma (DLBCL) and follicular lymphomas (FL). Mutations altering EZH2 tyrosine 641 (e.g., Y641C, Y641F, Y641N, Y641S, and Y641H) were reportedly observed in up to 22% of germinal center B-cell DLBCL and 7% of FL. Morin et al. Nat. Genetics 2010 February; 42(2):181-185. Mutations of alanine 677 (A677) and alanine 687 (A687) have also been reported. McCabe et al., Proc. Natl. Acad. Sci. USA 2012, 109:2989-2994; Majer et al. FEBS Letters 2012, 586:3448-3451. EZH2 activating mutations have been suggested to alter substrate specificity resulting in elevated levels of trimethylated H3K27 (H3K27me3).Accordingly, compounds that inhibit the activity of wild type and/or mutant forms of EZH2 may be of interest for the treatment of cancer.
SYNTHESIS

Steps
1 COUPLING, Ag2CO3
2 Alkylation, K2CO3
3 LiAlH4 REDUCTION
4 THIONYL CHLORIDE
5 N-Alkylation of Amides, t-BuOK
6 A GRIGNARD REACTION
7 AN ALKYLATION , METHYL IODIDE, t-BuOK
8 HYDROGENATION, DE BENZYLATION, PLATINUM OXIDE
9 LAST STEP separation by chiral preparative, SFC on (R,R) Whelk O1 column, TO GET PF 06821497
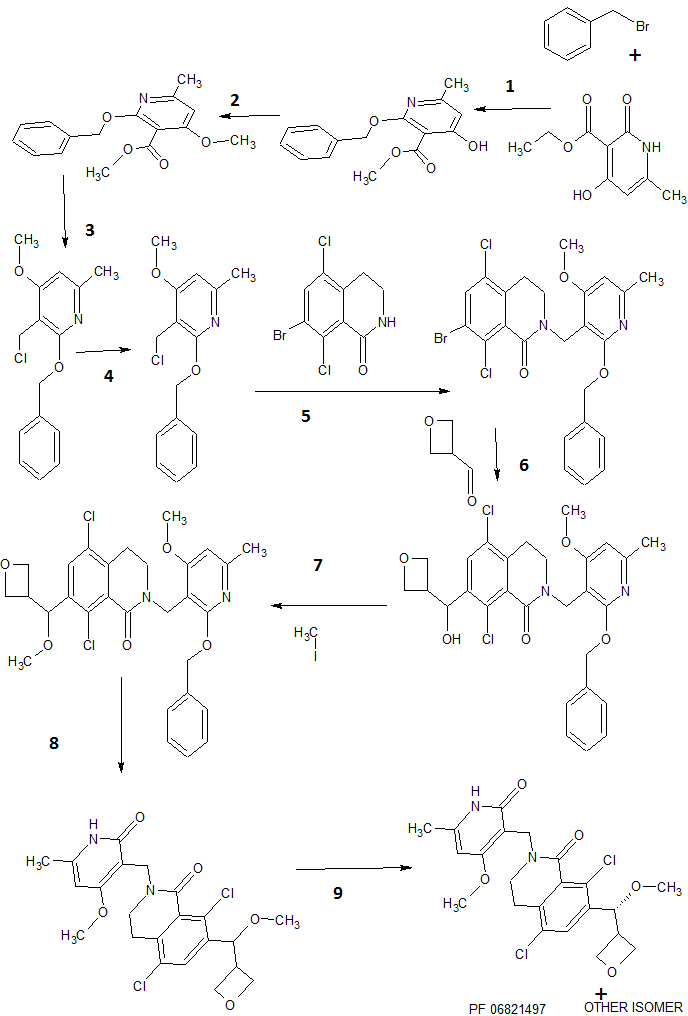
PATENT
US 20150361067

///////////////PF 06821497, 1844849-11-1, PFIZER, lymphoma, Pei-Pei Kung, @pfizer, #ACSSanFran, Michael Raymond Collins, Robert Steven Kania, Robert Arnold Kumpf, Pei-Pei Kung, Daniel Tyler Richter, Scott Channing Sutton, Martin James Wythes
Next up in #MEDI 1st time disclosures Pei-Pei Kung from @pfizer presenting a molecule designed to treat lymphoma #ACSSanFran
CO[C@H](c2cc(Cl)c3CCN(CC1=C(OC)C=C(C)NC1=O)C(=O)c3c2Cl)C4COC4
| CC1=CC(=C(C(=O)N1)CN2CCC3=C(C=C(C(=C3C2=O)Cl)C(C4COC4)OC)Cl)OC |
BLU 554
BLU 554
FGFR4 Inhibitor
N-[(3S,4S)-3-[[6-(2,6-Dichloro-3,5-dimethoxyphenyl)-2-quinazolinyl]amino]tetrahydro-2H-pyran-4-yl]-2-propenamide
N-[(3S,4S)-3-[[6-(2,6-Dichloro-3,5-dimethoxyphenyl)quinazolin-2-yl]amino]tetrahydro-2H-pyran-4-yl]acrylamide
- CAS No. 1707289-21-1
- Formula C24H24Cl2N4O4
- MolWeight 503.378
PHASE 1
BLU-554 is a potent fibroblast growth factor receptor 4 (FGFR4) inhibitor.
IC50 & Target: FGFR4[1]
InVitro: Fibroblast growth factor receptor 4 (FGFR-4) is a protein that in humans is encoded by the FGFR-4 gene. This protein is a member of the fibroblast growth factor receptor family, where amino acid sequence was highly conserved between members throughout evolution. FGFR family members 1-4 differ from one another in their ligand affinities and tissue distribution. A full-length representative protein consists of an extracellular region composed of three immunoglobulin-like domains, a single hydrophobic membrane-spanning segment and a cytoplasmic tyrosine kinase domain. The extracellular portion of the protein interacts with fibroblast growth factors, setting in motion a cascade of downstream signals, ultimately influencing mitogenesis and differentiation. The genomic organization of the FGFR-4 gene encompasses 18 exons. Although alternative splicing has been observed, there is no evidence that the C-terminal half of the Iglll domain of this protein varies between three alternate forms, as indicated for FGFR 1-3[1].
| Inventors | Neil Bifulco, Lucian V. Dipietro, Brian L. Hodous, Chandrasekhar V. MIDUTURU |
| Applicant | Blueprint Medicines Corporation |

Neil Bifulco
Senior Scientist at Blueprint Medicines

Chandra Miduturu
Senior Scientist at Blueprint Medicines
Fibroblast growth factor receptor 4 (FGFR-4) is a protein that in humans is encoded by the FGFR-4 gene. This protein is a member of the fibroblast growth factor receptor family, where amino acid sequence was highly conserved between members throughout evolution. FGFR family members 1-4 differ from one another in their ligand affinities and tissue distribution. A full-length representative protein consists of an extracellular region composed of three immunoglobulin-like domains, a single hydrophobic membrane-spanning segment and a cytoplasmic tyrosine kinase domain. The extracellular portion of the protein interacts with fibroblast growth factors, setting in motion a cascade of downstream signals, ultimately influencing mitogenesis and differentiation. The genomic organization of the FGFR-4 gene encompasses 18 exons. Although alternative splicing has been observed, there is no evidence that the C-terminal half of the Iglll domain of this protein varies between three alternate forms, as indicated for FGFR 1-3.
Ectopic mineralization, characterized by inappropriate calcium-phosphorus deposition in soft tissue, has been observed in rats treated with an FGFR-1 inhibitor (Brown, AP et al. (2005), Toxicol. Pathol., p. 449-455). This suggests that selective inhibition of FGFR-4 without inhibition of other isoforms of FGFR, including FGFR-1, may be desirable in order to avoid certain toxicities. FGFR-4 preferentially binds fibroblast growth factor 19 (FGF19) and has recently been associated with the progression of certain sarcomas, renal cell cancer, breast cancer, and liver cancer.
PATENT
PATENT
Synthetic Protocol 3

2-chloro-6-(2,6-dichloro-3,5-dimethoxyphenyl)quinazoline (described in WO 2014011900) can be substituted with an 1,2-mono-protected cycloalkyldiamine under various nucleophilic aromatic substitution reaction conditions using a base (such as diisopropylethylamine (DIPEA), DBU or NaHC03) in a polar solvent (such as dioxane, CH CN or NMP) or via a palladium-mediated Buchwald coupling reaction to provide the diamine- substituted quinazoline. The protecting group on the amine is removed to reveal the amine on the cycloalkane. The amine can be reacted with propiolic acid using amide coupling reaction conditions or reacted with acryloyl chloride to prepare the acrylamide. As shown below, Compounds 27, 32, 34, 36, and 40 were prepared using Synthetic Protocol 3.
Compound 40
Synthesis of N-((3S,4S)-3-((6-(2,6-dichloro-3,5-dimethoxyphenyl)quinazolin-2-yl)amino)tetrahydro-2H-pyran-4-yl)acrylamide

Step 1: Synthesis of N-((3S,4S)-4-azidotetrahydro-2H-pyran-3-yl)-6-(2,6-dichloro-3,5-dimethoxyphenyl)quinazolin-2-amine

(3S,4S)-4-azidotetrahydro-2H-pyran-3-amine, HC1 (0.200 g, 1.120 mmol) and 2-chloro-6-(2,6-dichloro-3,5-dimethoxyphenyl)quinazoline (0.318 g, 0.861 mmol) were taken up in NMP (2 ml) and sodium carbonate (0.217 g, 2.58 mmol) was added. The reaction was heated to 100 °C overnight. After cooling to ambient temperature the reaction was poured into 5ml of water and stirred for 30 min. The solid layer was filtered off and washed with water and further dried under high vacuum to give N-((3S,4S)-4-azidotetrahydro-2H-pyran-3-yl)-6-(2,6-dichloro-3,5-dimethoxyphenyl)quinazolin-2-amine (0.300 g, 0.631 mmol, 73.3 % yield). MS (ES+) C21H20CI2N6O3requires: 474, found: 475 [M + H]+.
Step 2: Synthesis of (3S,4S)-N3-(6-(2,6-dichloro-3,5-dimethoxyphenyl)quinazolin-2-yl)tetrahydro-2H-pyran-3,4-diamine

N-((3S,4S)-4-azidotetrahydro-2H-pyran-3-yl)-6-(2,6-dichloro-3,5-dimethoxyphenyl)quinazolin-2-amine (0.063 g, 0.133 mmol) was taken up in Methanol (7 ml) and EtOAc (7.00 ml), Pd-C (0.014 g, 0.133 mmol) was added and stirred under a ¾ balloon for 1 hour. After the reaction was completed, it was filtered through celite and the solvent removed. (3S,4S)-N3-(6-(2,6-dichloro-3,5-dimethoxyphenyl)quinazolin-2-yl)tetrahydro-2H-pyran-3,4-diamine (0.060 g, 0.134 mmol, 101 % yield) was recovered as a yellow solid, which was carried on without further purification. MS (ES+) C21H22CI2N4O3 requires: 448, found: 449 [M + H]+.
Step 3: Synthesis of N-((3S,4S)-3-((6-(2,6-dichloro-3,5-dimethoxyphenyl)quinazolin-2-yl)amino)tetrahydro-2H-pyran-4-yl)acrylamide

(3S,4S)-N3-(6-(2,6-dichloro-3,5-dimethoxyphenyl)quinazolin-2-yl)tetrahydro-2H-pyran-3,4-diamine (0.060 g, 0.134 mmol) was taken up in CH2CI2 (2 ml) and cooled to 0 °C, followed by addition of DIEA (0.023 ml, 0.134 mmol) and then acryloyl chloride (0.012 ml, 0.147 mmol) slowly. The reaction was stirred at 0 °C for 30 minutes, then the mixture was loaded directly onto silica and purified by flash chromotography using 0-10% CH2Cl2/MeOH. N-((3S,4S)-3-((6-(2,6-dichloro-3,5-dimethoxyphenyl)quinazolin-2-yl)amino)tetrahydro-2H-pyran-4-yl)acrylamide (0.041 g, 0.081 mmol, 61% yield) was recovered as an off white solid. MS (ES+) C24H24CI2N4O4 requires: 502, found: 503 [M + H]+.
References on BLU-554

Jeff Albers
Chief Executive Officer at Blueprint Medicines

Marion Dorsch
Chief Scientific Officer at Blueprint Medicines

Chris De Savi
Director of Medicinal Chemistry at Blueprint Medicines

Blueprint Medicines is developing a new generation of highly selective and potent kinase therapies to dramatically improve the lives of patients with genomically defined diseases. Our approach is rooted in a deep understanding of the genetic blueprint of cancer and other diseases driven by the abnormal activation of kinases. Our ability to identify novel drivers of disease, coupled with our proprietary library of novel and diverse chemical compounds, uniquely enables us to craft kinase therapies against new and difficult-to-drug targets. We are boldly advancing a deep pipeline of highly targeted therapies against previously unaddressed drivers of disease. By focusing on genomically defined subsets of patients, we believe we can identify the people most likely to respond to our therapies, resulting in a more efficient clinical development path with a greater likelihood of success and better outcomes for patients. We see a substantial opportunity in kinase drug discovery and development to deliver breakthrough medicines that allow patients to live longer with better quality of life and prevent recurrences of disease. Kinases are involved in many hallmarks of tumor biology and are proven cancer drug targets. Currently approved drugs focus on less than 5 percent of known kinases, and the function of most kinases is unknown. Led by a team of industry innovators with a track record of bringing life-changing drugs to market, we believe Blueprint Medicines has the experience and expertise to deliver on the tremendous untapped potential of kinase therapies to improve patients’ lives. We don’t think in small steps. We think in giant leaps. We are driven by the pursuit of new ideas, new innovations, and new ways of thinking.
Specialties
Oncology drug discovery, Oncology, Genomically defined diseases, Rare diseases, Transformative drugs, Robust drug pipeline, Systemic mastocytosis, Hepatocellular carcinoma, Orphan drugs, Small molecules, Kinase inhibitor, Personalized medicine, and Novel cancer therapies
//////////BLU 554, FGFR4 Inhibitor, Chandra Miduturu, @BlueprintMeds, advanced heptocellular carcinoma, #ACSSanFran, PHASE 1, Neil Bifulco, Lucian V. Dipietro, Brian L. Hodous, Chandrasekhar V. MIDUTURU, BLUEPRINT,
Now Chandra Miduturu of @BlueprintMeds is speaking in #MEDI 1st time disclosures about of advanced heptocellular carcinoma #ACSSanFran
 DRUG APPROVALS BY DR ANTHONY MELVIN CRASTO
.....
DRUG APPROVALS BY DR ANTHONY MELVIN CRASTO
.....















